# 1. Request free access at https://listoagriculture.com/products
# 2. Follow the installation instructions sent to your emailIntroduction to Mandala
Getting started with agricultural field trial analysis
Overview
The mandala package provides tools for analyzing agricultural field trial data using mixed models. This tutorial introduces the basic concepts and functionality of the package.
- Installing and loading Mandala
- Understanding the key features
- Basic model structure for field trials
- Running a simple analysis
- Visualizing results
Installation
To install Mandala, request free access at Listo Agriculture. You will receive an email with instructions to install the package from the Listo repository.
Loading Required Packages
library(mandala)
library(dplyr)
library(ggplot2)Key Features
The mandala package offers:
- Mixed Model Analysis: Fit linear mixed models optimized for agricultural field trials
- Spatial Analysis: AR1 correlation structures and P-spline smoothing (
pspline2D) with variogram diagnostics - Genomic Prediction: GBLUP with relationship matrices via
GM()and cross-validation withmandala_gp_cv() - Factor-Analytic G×E: Model complex genotype-by-environment interactions with
FA(), biplots, and reliability estimates - Variance Component Estimation: Extract and interpret genetic and environmental variances
- Heritability: Unified estimation via
h2_estimates()with Cullis and Piepho methods - Prediction: Generate genotype BLUEs from fixed-genotype fits and BLUPs from random-genotype fits
Basic Model Structure
In agricultural field trials, we typically have:
| Effect Type | Examples | Purpose |
|---|---|---|
| Fixed | Treatments, checks, genotypes when estimating BLUEs | Effects we want to estimate directly |
| Random | Blocks, incomplete blocks, genotypes when predicting BLUPs | Effects we want to predict or use for variance partitioning |
| Spatial | Row, column correlations | Account for field heterogeneity |
Whether genotype is fixed or random changes the interpretation of the results: fixed-genotype fits support genotype BLUEs, while random-genotype fits support genotype BLUPs.
A typical model might be:
\[ y_{ijk} = \mu + \tau_i + b_j + \epsilon_{ijk} \]
Where:
- \(y_{ijk}\) is the observed response (e.g., yield)
- \(\mu\) is the overall mean
- \(\tau_i\) is the fixed treatment effect (genotype)
- \(b_j \sim N(0, \sigma^2_b)\) is the random block effect
- \(\epsilon_{ijk} \sim N(0, \sigma^2_e)\) is the residual error
Example: Simple Analysis
Let’s create a simple example dataset to demonstrate the workflow:
# Create example field trial data
set.seed(123)
n_genotypes <- 10
n_blocks <- 4
n_obs <- n_genotypes * n_blocks
example_data <- data.frame(
genotype = factor(rep(1:n_genotypes, each = n_blocks)),
block = factor(rep(1:n_blocks, times = n_genotypes)),
yield = rnorm(n_obs, mean = 100, sd = 10)
)
# Add treatment effects
treatment_effects <- rnorm(n_genotypes, mean = 0, sd = 5)
example_data$yield <- example_data$yield +
treatment_effects[as.numeric(example_data$genotype)]
# Add block effects
block_effects <- rnorm(n_blocks, mean = 0, sd = 3)
example_data$yield <- example_data$yield +
block_effects[as.numeric(example_data$block)]
head(example_data) genotype block yield
1 1 1 91.68166
2 1 2 94.13905
3 1 3 111.98494
4 1 4 101.33736
5 2 1 101.01325
6 2 2 116.02542Visualizing the Data
Before fitting any model, it’s important to explore your data:
# Plot yield by genotype
ggplot(example_data, aes(x = genotype, y = yield, color = block)) +
geom_point(size = 3, alpha = 0.6) +
theme_minimal() +
labs(title = "Yield by Genotype and Block",
x = "Genotype", y = "Yield")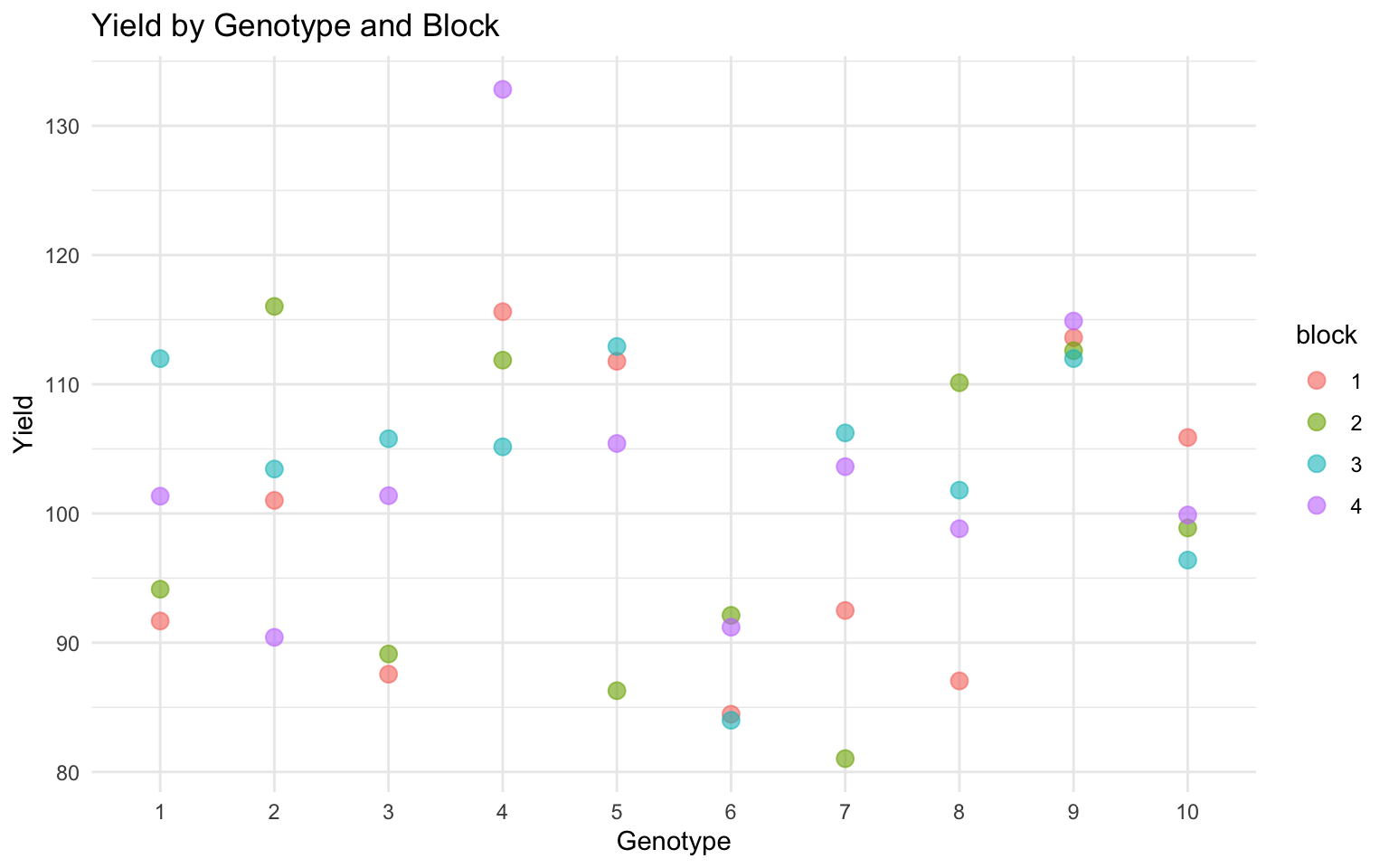
# Summary statistics by genotype
example_data %>%
group_by(genotype) %>%
summarise(
mean_yield = mean(yield),
sd_yield = sd(yield),
n = n()
) %>%
head()# A tibble: 6 × 4
genotype mean_yield sd_yield n
<fct> <dbl> <dbl> <int>
1 1 99.8 9.11 4
2 2 103. 10.5 4
3 3 96.0 9.00 4
4 4 116. 11.8 4
5 5 104. 12.3 4
6 6 87.9 4.31 4Basic Model Fitting
Mandala uses a clear formula interface:
fixed = yield ~ genotype— fixed effects formularandom = ~ block— random effects formula
data = example_data— your data frame
# Fit a basic mixed model
model <- mandala(
fixed = yield ~ genotype,
random = ~ block,
data = example_data
)Initial data rows: 40
Final data rows after NA handling: 40 --- Starting EM Warmup ---
EM Warmup: starting theta:
block 50.97 R.sigma2 63.71
EM iter 1: logLik=-117.9496; theta: block 33.32 R.sigma2 57.63
EM iter 2 : logLik decreased or failed. Stopping EM.
EM Warmup: ending theta:
block 33.32 R.sigma2 57.63
--- EM Warmup Complete. Starting AI-REML ---Starting AI-REML logLik = -117.9753
Iter LogLik Sigma2 DF wall Restrained
1 -116.8878 71.464 30 15:02:36 ( 0 restrained)
2 -116.5873 88.615 30 15:02:36 ( 0 restrained)
3 -116.5550 93.559 30 15:02:36 ( 0 restrained)
4 -116.4094 91.300 30 15:02:36 ( 0 restrained)
5 -116.2541 86.964 30 15:02:36 ( 0 restrained)
6 -116.1432 83.577 30 15:02:36 ( 0 restrained)
7 -116.0649 82.011 30 15:02:36 ( 0 restrained)
8 -116.0024 81.868 30 15:02:36 ( 0 restrained)
9 -115.9508 82.382 30 15:02:36 ( 0 restrained)
10 -115.9084 82.956 30 15:02:36 ( 0 restrained)
11 -115.8729 83.315 30 15:02:36 ( 0 restrained)
12 -115.8428 83.432 30 15:02:36 ( 0 restrained)
13 -115.8175 83.399 30 15:02:36 ( 0 restrained)
14 -115.7967 83.316 30 15:02:36 ( 0 restrained)
15 -115.7800 83.248 30 15:02:36 ( 0 restrained)
16 -115.7667 83.214 30 15:02:36 ( 0 restrained)
17 -115.7561 83.209 30 15:02:36 ( 0 restrained)
18 -115.7477 83.218 30 15:02:36 ( 0 restrained)
19 -115.7411 83.229 30 15:02:36 ( 0 restrained)
20 -115.7357 83.237 30 15:02:36 ( 0 restrained)
21 -115.7315 83.239 30 15:02:36 ( 0 restrained)
22 -115.7281 83.239 30 15:02:36 ( 0 restrained)
23 -115.7254 83.237 30 15:02:36 ( 0 restrained)
Froze block at 1.000006e-05
24 -115.7233 83.236 30 15:02:36 ( 1 restrained)
25 -115.7136 83.235 30 15:02:36 ( 1 restrained)
26 -115.7136 83.235 30 15:02:36 ( 1 restrained)
27 -115.7136 83.235 30 15:02:36 ( 1 restrained)
Converged at iter 27 by tolerance(s): delta_theta=0.00e+00, delta_loglik=0.00e+00
Main AI-REML Loop finished. Combining results.# View results
summary(model)Model statement:
mandala(fixed = yield ~ genotype, random = ~block, data = example_data)
Variance Components:
component estimate std.error z.ratio bound %ch
block 1.000006e-05 7.163753 1.395924e-06 B NA
R.sigma2 8.323505e+01 22.653751 3.674228e+00 P NA
Fixed Effects (BLUEs) [first 5]:
effect estimate std.error z.ratio
(Intercept) 99.785670 4.561640 21.8749564
genotype2 2.938138 6.451144 0.4554445
genotype3 -3.821318 6.451144 -0.5923474
genotype4 16.578287 6.451144 2.5698213
genotype5 4.316508 6.451144 0.6691074
Converged: TRUE | Iterations: 27
Random Effects (BLUPs) [first 5]:
random level estimate std.error z.ratio
block 1 -2.950838e-06 0.003162285 -0.0009331348
block 2 -2.825363e-06 0.003162285 -0.0008934560
block 3 2.883142e-06 0.003162285 0.0009117274
block 4 2.893107e-06 0.003162285 0.0009148786
logLik: -115.714 AIC: 235.427 BIC: 238.230 logLik_Trunc: -88.145# View variance components
var_comps <- model$varcomp
print(var_comps) component estimate std.error z.ratio bound %ch
1 block 1.000006e-05 7.163753 1.395924e-06 B NA
2 R.sigma2 8.323505e+01 22.653751 3.674228e+00 P NA# Get adjusted genotype means (BLUEs when genotype is fixed)
predictions <- mandala_predict(model, classify_term = "genotype")
Notes:
- The predictions are obtained by averaging across the hypertable
calculated from model terms constructed solely from factors in
the averaging and classify sets.
- The simple averaging set: (none)head(predictions) genotype predicted_value std_error
1 1 99.78567 4.56164
2 10 100.25996 4.56166
3 2 102.72381 4.56166
4 3 95.96435 4.56166
5 4 116.36396 4.56166
6 5 104.10218 4.56166Model Output Interpretation
When you run summary(model), you’ll typically see:
- Variance Components Table
- Block variance (σ²_b)
- Residual variance (σ²_e)
- Fixed Effects
- Intercept and genotype-effect coefficients
- Standard errors
- Model Fit Statistics
- Log-likelihood
- AIC/BIC
To obtain adjusted genotype means from this fixed-genotype model, use mandala_predict(model, classify_term = "genotype").
Next Steps
Now that you understand the basics, continue with:
- Tutorial 02: Working with Example Datasets — Real-world experimental designs
- Tutorial 03: Comparison with Sommer — Genomic prediction and quantitative genetics
- Tutorial 04: Comparison with lme4 — General mixed model context
- Tutorial 05: Comparison with SpATS — P-spline spatial analysis
References
- Mandala: https://listoagriculture.com/products
- Piepho, H.P., et al. (2003). BLUP for phenotypic selection in plant breeding
- Best practices for field trial analysis
Session Information
sessionInfo()R version 4.5.2 (2025-10-31)
Platform: aarch64-apple-darwin20
Running under: macOS Tahoe 26.1
Matrix products: default
BLAS: /System/Library/Frameworks/Accelerate.framework/Versions/A/Frameworks/vecLib.framework/Versions/A/libBLAS.dylib
LAPACK: /Library/Frameworks/R.framework/Versions/4.5-arm64/Resources/lib/libRlapack.dylib; LAPACK version 3.12.1
locale:
[1] C.UTF-8/C.UTF-8/C.UTF-8/C/C.UTF-8/C.UTF-8
time zone: America/New_York
tzcode source: internal
attached base packages:
[1] stats graphics grDevices utils datasets methods base
other attached packages:
[1] ggplot2_4.0.0 dplyr_1.1.4 mandala_1.1.0
loaded via a namespace (and not attached):
[1] vctrs_0.6.5 cli_3.6.5 knitr_1.50 rlang_1.1.6
[5] xfun_0.54 generics_0.1.4 S7_0.2.0 splines2_0.5.4
[9] jsonlite_2.0.0 labeling_0.4.3 glue_1.8.0 htmltools_0.5.8.1
[13] scales_1.4.0 rmarkdown_2.30 grid_4.5.2 evaluate_1.0.5
[17] tibble_3.3.0 fastmap_1.2.0 yaml_2.3.10 lifecycle_1.0.4
[21] compiler_4.5.2 codetools_0.2-20 RColorBrewer_1.1-3 htmlwidgets_1.6.4
[25] Rcpp_1.1.0 pkgconfig_2.0.3 farver_2.1.2 lattice_0.22-7
[29] digest_0.6.37 R6_2.6.1 utf8_1.2.6 tidyselect_1.2.1
[33] pillar_1.11.1 magrittr_2.0.4 Matrix_1.7-4 withr_3.0.2
[37] gtable_0.3.6 tools_4.5.2