library(mandala)
library(lme4)
library(dplyr)
library(ggplot2)
get_mandala_vc <- function(varcomp, term) {
hits <- which(varcomp$component == term)
if (length(hits) != 1L) {
stop("Expected exactly one Mandala variance component for term: ", term)
}
varcomp$estimate[hits]
}
get_lme4_vc <- function(varcomp, grp) {
hits <- which(varcomp$grp == grp)
if (length(hits) != 1L) {
stop("Expected exactly one lme4 variance component for group: ", grp)
}
varcomp$vcov[hits]
}Comparison: Mandala vs lme4
Field trial analysis vs general-purpose mixed models
Overview
This tutorial compares the mandala package with lme4, the most widely-used R package for mixed models. Where both packages support the same linear mixed model directly, the examples below fit the same structure in both packages. Where Mandala extends beyond base lme4 with field-trial-specific spatial terms and postprocessing, those features are shown as workflow extensions rather than as one-to-one model matches.
- How the same RCBD and nested mixed models look in Mandala and
lme4 - Which outputs are directly comparable across the two packages
- Where Mandala adds field-trial-specific helpers on top of a standard mixed-model workflow
- When differences reflect model targets or workflow scope rather than a disagreement in results
Background
| Package | Primary Focus | Best For |
|---|---|---|
| lme4 | General-purpose | Any discipline |
| Mandala | Agricultural trials | Field experiments |
Loading Packages
Example 1: Basic RCBD
Data Setup
set.seed(321)
n_genotypes <- 12
n_blocks <- 5
rcbd_data <- expand.grid(
genotype = factor(paste0("G", sprintf("%02d", 1:n_genotypes))),
block = factor(paste0("B", 1:n_blocks))
)
rcbd_data$yield <- 4500 +
rnorm(n_genotypes, 0, 20)[as.numeric(rcbd_data$genotype)] +
rnorm(n_blocks, 0, 10)[as.numeric(rcbd_data$block)] +
rnorm(nrow(rcbd_data), 0, 15)
head(rcbd_data) genotype block yield
1 G01 B1 4542.286
2 G02 B1 4496.306
3 G03 B1 4503.019
4 G04 B1 4513.249
5 G05 B1 4497.798
6 G06 B1 4522.072Syntax Comparison
Mandala:
mandala(
fixed = yield ~ genotype,
random = ~ block,
data = rcbd_data
)lme4:
lmer(
yield ~ genotype + (1|block),
data = rcbd_data
)Mandala RCBD
mandala_rcbd <- mandala(
fixed = yield ~ genotype,
random = ~ block,
data = rcbd_data,
verbose = FALSE
)
mandala_rcbd_vc <- mandala_rcbd$varcomp
mandala_rcbd_coef <- mandala_rcbd$BLUEs %>%
transmute(effect, mandala_estimate = estimate)
print(mandala_rcbd_vc) component estimate std.error z.ratio bound %ch
1 block 129.1804 102.67498 1.258149 P NA
2 R.sigma2 215.5579 40.79218 5.284295 P NAhead(mandala_rcbd_coef) effect mandala_estimate
(Intercept) (Intercept) 4529.89386
genotypeG02 genotypeG02 -47.16105
genotypeG03 genotypeG03 -33.39576
genotypeG04 genotypeG04 -19.82680
genotypeG05 genotypeG05 -28.04273
genotypeG06 genotypeG06 -16.66351lme4 RCBD
lme4_model <- lmer(
yield ~ genotype + (1|block),
data = rcbd_data,
REML = TRUE
)
lme4_rcbd_vc <- as.data.frame(VarCorr(lme4_model))
lme4_rcbd_coef <- tibble(
effect = names(fixef(lme4_model)),
lme4_estimate = unname(fixef(lme4_model))
)
print(lme4_rcbd_vc) grp var1 var2 vcov sdcor
1 block (Intercept) <NA> 125.5453 11.20470
2 Residual <NA> <NA> 221.6143 14.88671head(lme4_rcbd_coef)# A tibble: 6 × 2
effect lme4_estimate
<chr> <dbl>
1 (Intercept) 4530.
2 genotypeG02 -47.2
3 genotypeG03 -33.4
4 genotypeG04 -19.8
5 genotypeG05 -28.0
6 genotypeG06 -16.7RCBD Comparison
rcbd_coef_compare <- inner_join(
mandala_rcbd_coef,
lme4_rcbd_coef,
by = "effect"
)
head(rcbd_coef_compare) effect mandala_estimate lme4_estimate
1 (Intercept) 4529.89386 4529.90137
2 genotypeG02 -47.16105 -47.16590
3 genotypeG03 -33.39576 -33.40059
4 genotypeG04 -19.82680 -19.83162
5 genotypeG05 -28.04273 -28.04756
6 genotypeG06 -16.66351 -16.66832tibble(
package = c("Mandala", "lme4"),
block_variance = c(get_mandala_vc(mandala_rcbd_vc, "block"),
get_lme4_vc(lme4_rcbd_vc, "block")),
residual_variance = c(get_mandala_vc(mandala_rcbd_vc, "R.sigma2"),
get_lme4_vc(lme4_rcbd_vc, "Residual"))
)# A tibble: 2 × 3
package block_variance residual_variance
<chr> <dbl> <dbl>
1 Mandala 129. 216.
2 lme4 126. 222.What Differs Here
Both packages are fitting the same fixed-genotype RCBD model in this section, so the coefficient and variance-component estimates should be close. The main difference is workflow: Mandala exposes a field-trial-oriented fixed-effect solution table directly through model$BLUEs, while lme4 gives the same fixed-effect information through fixef() and the variance components through VarCorr().
Example 2: Nested Design
Data Setup
set.seed(654)
n_genotypes <- 30
n_complete <- 3
nested_data <- expand.grid(
genotype = factor(paste0("G", sprintf("%02d", 1:n_genotypes))),
complete_block = factor(1:n_complete)
)
nested_data$incomplete_block <- factor(
paste0(nested_data$complete_block, ".",
rep(1:5, length.out = nrow(nested_data)))
)
nested_data$yield <- 5500 +
rnorm(n_genotypes, 0, 30)[as.numeric(nested_data$genotype)] +
rnorm(n_complete, 0, 15)[as.numeric(nested_data$complete_block)] +
rnorm(nrow(nested_data), 0, 20)Mandala Nested Design
mandala_nested <- mandala(
fixed = yield ~ genotype,
random = ~ complete_block + complete_block:incomplete_block,
data = nested_data,
verbose = FALSE
)
mandala_nested_vc <- mandala_nested$varcomp
print(mandala_nested_vc) component estimate std.error z.ratio bound %ch
1 complete_block 1.000090e-05 16.69320 5.991001e-07 B NA
2 complete_block:incomplete_block 2.448528e+01 38.65217 6.334773e-01 P NA
3 R.sigma2 3.010141e+02 60.20275 5.000007e+00 P NAlme4 Nested Design
lme4_nested <- lmer(
yield ~ genotype + (1|complete_block) + (1|complete_block:incomplete_block),
data = nested_data,
REML = TRUE
)
lme4_nested_vc <- as.data.frame(VarCorr(lme4_nested))
print(lme4_nested_vc) grp var1 var2 vcov sdcor
1 complete_block:incomplete_block (Intercept) <NA> 24.4848159 4.9482134
2 complete_block (Intercept) <NA> 0.7170808 0.8468062
3 Residual <NA> <NA> 301.0144193 17.3497671Nested Design Comparison
tibble(
package = c("Mandala", "lme4"),
complete_block_variance = c(get_mandala_vc(mandala_nested_vc, "complete_block"),
get_lme4_vc(lme4_nested_vc, "complete_block")),
incomplete_block_variance = c(get_mandala_vc(mandala_nested_vc, "complete_block:incomplete_block"),
get_lme4_vc(lme4_nested_vc, "complete_block:incomplete_block")),
residual_variance = c(get_mandala_vc(mandala_nested_vc, "R.sigma2"),
get_lme4_vc(lme4_nested_vc, "Residual"))
)# A tibble: 2 × 4
package complete_block_variance incomplete_block_variance residual_variance
<chr> <dbl> <dbl> <dbl>
1 Mandala 0.0000100 24.5 301.
2 lme4 0.717 24.5 301.What Differs Here
This example stays within the overlap between the two packages, so the nested-block variance decomposition is directly comparable. When a variance component is near zero, the two optimizers may report slightly different boundary behavior, but the practical conclusion is the same if both fits place little support on that component.
Diagnostics
Both packages support diagnostics, but they package them differently: lme4 exposes fitted values and residuals for general R plotting workflows, while Mandala bundles common field-trial diagnostics into dedicated helpers.
lme4 Diagnostics
diag_data <- data.frame(
Fitted = fitted(lme4_model),
Residuals = residuals(lme4_model)
)
p1 <- ggplot(diag_data, aes(x = Fitted, y = Residuals)) +
geom_point(alpha = 0.6, color = "#5D4E6D") +
geom_hline(yintercept = 0, linetype = "dashed", color = "#9B5B5B") +
geom_smooth(se = FALSE, color = "#8B9A6D") +
theme_minimal() +
labs(title = "Residuals vs Fitted")
p2 <- ggplot(diag_data, aes(sample = Residuals)) +
stat_qq(color = "#5D4E6D") +
stat_qq_line(color = "#9B5B5B") +
theme_minimal() +
labs(title = "Normal Q-Q Plot")
gridExtra::grid.arrange(p1, p2, ncol = 2)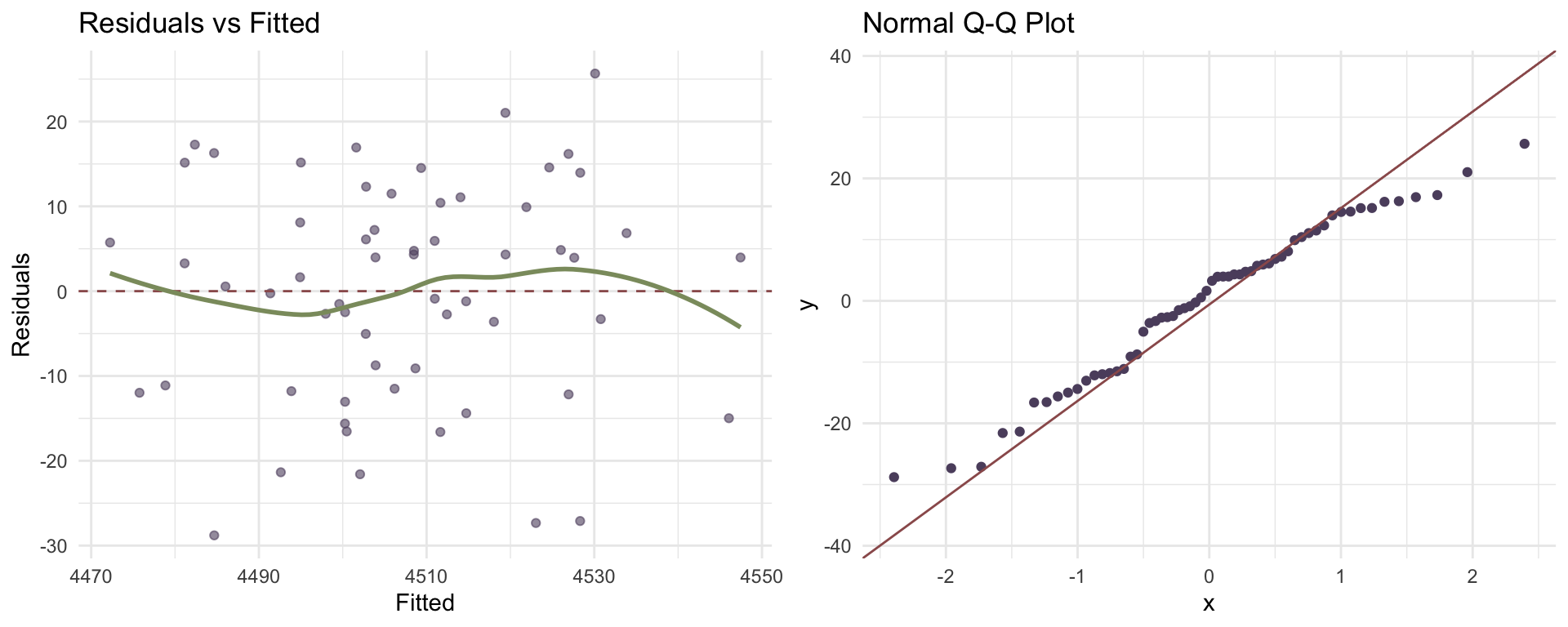
Mandala Diagnostics
# Built-in diagnostic plots for the Mandala RCBD fit
mandala_diagnostic_plots(mandala_rcbd, response = "yield")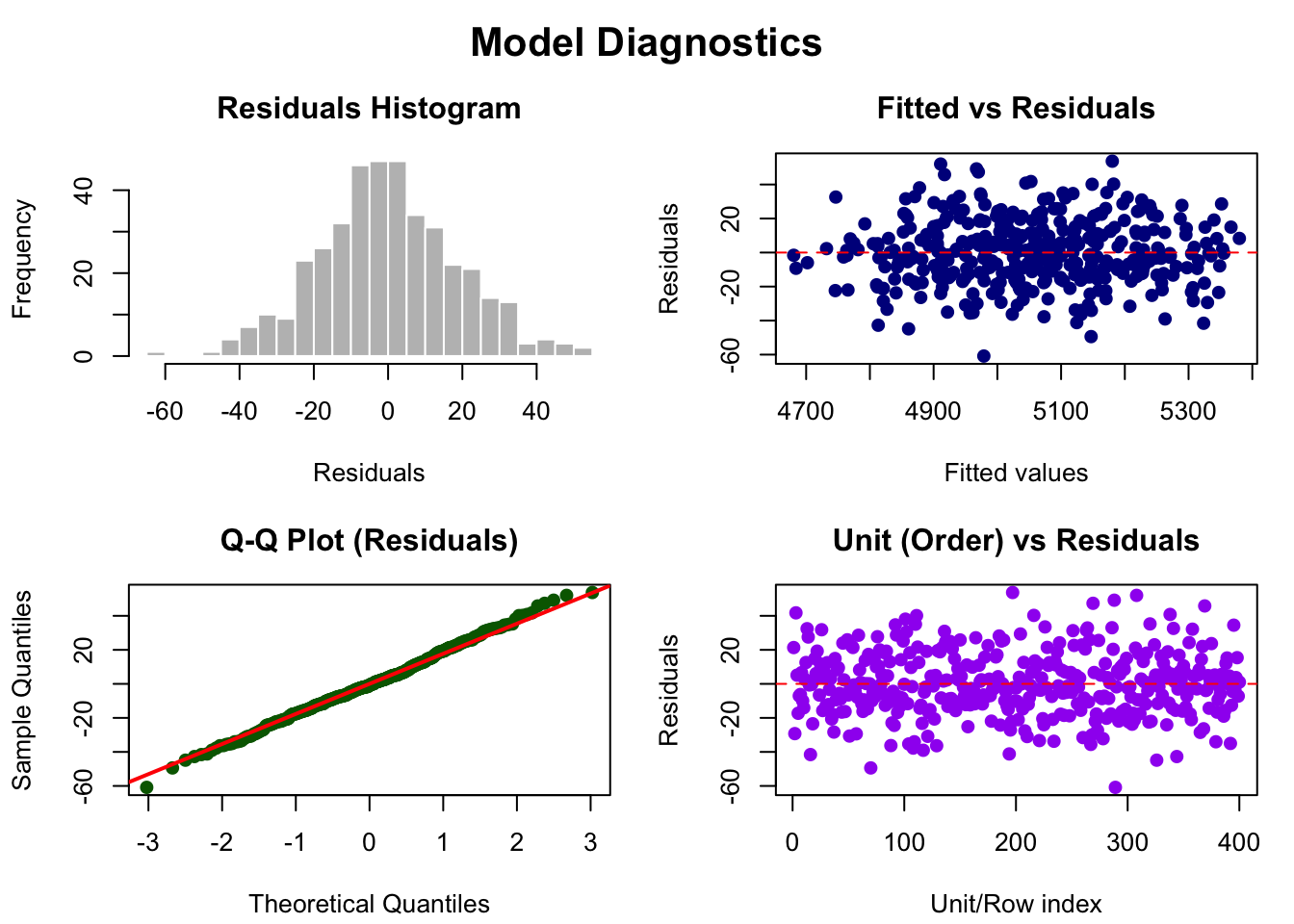
# Includes:
# - Residuals vs fitted
# - Q-Q plot
# - Histogram of residuals
# - Observed vs predictedKey Differences
| Task | Mandala | lme4 |
|---|---|---|
| Random intercept | random = ~ block |
(1\|block) |
| Variance components | $varcomp |
VarCorr() |
| Fixed and random genotype outputs | $BLUEs, mandala_predict() |
fixef(), ranef(), manual adjusted means |
| Heritability | Built-in h2_estimates() |
Manual calculation |
| AR1 spatial | ar1(...) inside random = ~ ... |
Not in base lme4 |
| P-spline spatial | pspline2D(...) inside random = ~ ... |
Not in base lme4 |
| Variograms | mandala_variogram() |
Manual or other packages |
| Diagnostics | mandala_diagnostic_plots() |
Fitted/residual outputs + general plotting |
| Genomic models | GM() |
Not in base lme4 |
When to Use Each
Use Mandala for:
- Agricultural field trials
- Spatial correlation modeling (AR1, P-splines)
- Variogram diagnostics
- Quick heritability estimates
- Genomic prediction (GBLUP)
- Opinionated field-trial result tables and postprocessing
Use lme4 for:
- General mixed models outside agriculture
- Complex random effect structures
- Generalized linear mixed models (GLMMs)
- Large documentation and community support
Session Information
sessionInfo()R version 4.5.2 (2025-10-31)
Platform: aarch64-apple-darwin20
Running under: macOS Tahoe 26.1
Matrix products: default
BLAS: /System/Library/Frameworks/Accelerate.framework/Versions/A/Frameworks/vecLib.framework/Versions/A/libBLAS.dylib
LAPACK: /Library/Frameworks/R.framework/Versions/4.5-arm64/Resources/lib/libRlapack.dylib; LAPACK version 3.12.1
locale:
[1] C.UTF-8/C.UTF-8/C.UTF-8/C/C.UTF-8/C.UTF-8
time zone: America/New_York
tzcode source: internal
attached base packages:
[1] stats graphics grDevices utils datasets methods base
other attached packages:
[1] ggplot2_4.0.0 dplyr_1.1.4 lme4_1.1-37 Matrix_1.7-4 mandala_1.1.0
loaded via a namespace (and not attached):
[1] gtable_0.3.6 jsonlite_2.0.0 compiler_4.5.2 tidyselect_1.2.1
[5] Rcpp_1.1.0 splines2_0.5.4 gridExtra_2.3 scales_1.4.0
[9] splines_4.5.2 boot_1.3-32 yaml_2.3.10 fastmap_1.2.0
[13] lattice_0.22-7 R6_2.6.1 labeling_0.4.3 generics_0.1.4
[17] knitr_1.50 rbibutils_2.3 htmlwidgets_1.6.4 MASS_7.3-65
[21] tibble_3.3.0 nloptr_2.2.1 minqa_1.2.8 RColorBrewer_1.1-3
[25] pillar_1.11.1 rlang_1.1.6 utf8_1.2.6 xfun_0.54
[29] S7_0.2.0 cli_3.6.5 mgcv_1.9-3 withr_3.0.2
[33] magrittr_2.0.4 Rdpack_2.6.4 digest_0.6.37 grid_4.5.2
[37] lifecycle_1.0.4 nlme_3.1-168 reformulas_0.4.2 vctrs_0.6.5
[41] evaluate_1.0.5 glue_1.8.0 farver_2.1.2 rmarkdown_2.30
[45] tools_4.5.2 pkgconfig_2.0.3 htmltools_0.5.8.1